About the Murine4Covid database
(under construction. 如您有感兴趣的其他基因,请联系页面底部邮箱,我们会尽快更新。)
The pandemic of Corona Virus Disease 2019 (COVID-19), a respiratory disease caused by a novel severe acute respiratory syndrome coronavirus (SARS-CoV-2), is causing substantial morbidity and mortality. Along with the respiratory symptoms, underlying diseases in senior patients such as diabetes, hypertension and coronary heart disease are the most common comorbidities, which cause more sever outcomes, even death. During the attach and entry of SARS-CoV-2, the key protein involved is the Angiotensin I Converting Enzyme 2 (ACE2) located on the membrane of host cells. Here, we aim to curate an expression profile of Ace2 and other COVID-19 related genes across the available diabetes murine strains. Based on strictly manual curation and bioinformatics analysis of the publicly deposited expression datasets, Ace2 and other potential proteins such as Furin, Tmprss2, Ang, Ang2 were examined. We found that Ace2 expression is rather ubiquitous in three diabetes prone strains (db/db, ob/ob and DIO). With the most abundant datasets presence, liver shows a medium Ace2 level compared with lung, pancreatic islet, brain and even T cell. Age is a critical factor for Ace2 expression in db/db than the other two strains. Other four host genes except Ace2 showed correlation to each other at varied levels. To accelerate the researches of the interaction between COVID-19 and the underlying diseases, transcriptomics database of Murine4Covid (www.geneureka.org/Murine4Covid) will facilitate the design of research on COVID-19 and underlying diseases.
Please cite:
Transcriptomics Curation of SARS-CoV-2 Related Host Genes in Mice with COVID-19 Comorbidity: A Pilot Study
Kunkai Su#, Xin Huang#, Kaijin Xu, Weibo Du, Danhua Zhu, Meifang Yang, Wenji Yuan, Lanjuan Li*
Infectious Microbes & Diseases: April 17, 2020 - Volume Latest Articles - Issue -
doi: 10.1097/IM9.0000000000000025
Example: Ace2 profile for diabetes mouse
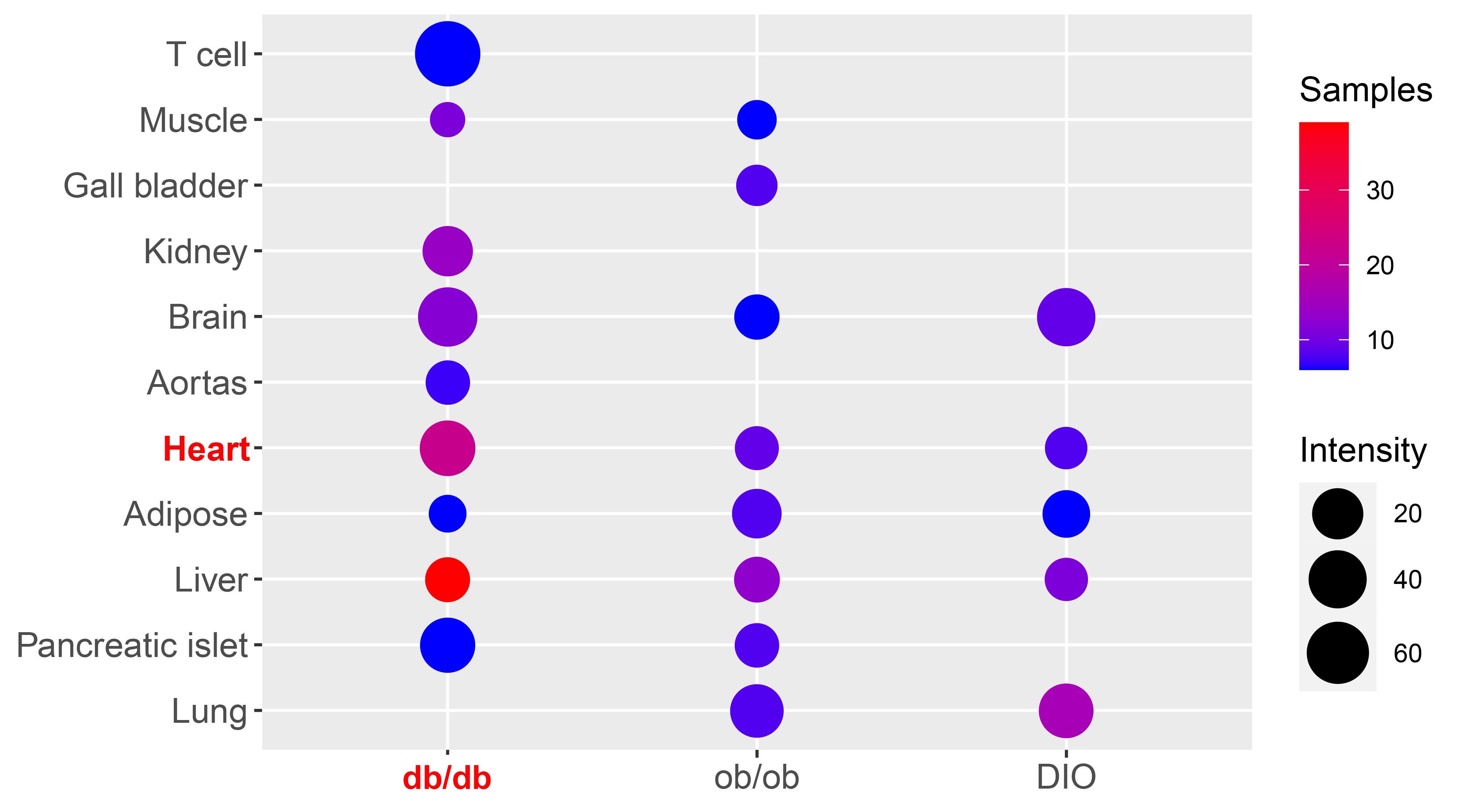
Summary of Ace2 expression in mouse tissues based on publicly available transcriptomics datasets. Different sizes of circles represent normalized expression levels of Ace2, and scaled colors of circles represent number of samples curated in certain tissue of strain. A consistent expression in the lung, pancreatic islet, liver, adipose, heart, aortas, brain, kidney, gall bladder, muscle and T cells was observed across all datasets. Blank in situ represents no qualified dataset available. Intensity is normalized and shown as to /(103 intensity of geometric mean of Gapdh and Actb).
Other gene and/or comabidity
Data in processing.
More specific target genes and murine strains will be added into the portfolio on request.